Table of Contents
Getting Started: The Cohead Salmon
We will build a simple knowledge base detailing the habitat requirements for a fictional fish, the Cohead salmon, at a geographically defined stream segment, or “reach”. Trainees should use this description of Cohead salmon biology/ecology as a tutorial exercise and should not consider it to be biologically accurate.
We recommend that you approach the development of your knowledge base by conceptualizing and building the outcome, or dependency network topic(s) that represent the broadest descriptions of the system. For the example of the Cohead salmon, we will build a dependency network for habitat quality that asks if Cohead can spawn and rear young in a particular stream segment. We will then refine this simple early model of the system by making each sub-process more explicit, a process called “building down”. Once one has built down to the level of elemental data (e.g., gravel size in the stream bed), then the prototype model can be considered complete.
NetWeaver Developer Toolbars
First, a quick introduction to the toolbars in NetWeaver Developer. The three main toolbars are organized by type of activity: application and knowledge base activities, network navigation, and network editing.
The First Network
Cohead salmon live in the ocean, but swim upstream to lay their eggs (spawn). For Cohead populations to be viable in a stream segment, conditions must permit the Cohead to spawn, and the young must be able to grow and swim to the sea (juvenile rearing).
We begin by organizing our knowledge base at the top (coarse description) level: Cohead Habitat. We will then build a dependency network that depicts the factors associated with assessing Cohead habitat quality by specifying the logical relationship of spawning and juvenile rearing habitats. In essence, the dependency network is a graphical depiction of a rule or set of rules.
Open NetWeaver Developer, then choose New from the File menu or click on the new project button  from the knowledge base toolbar. This creates a new project within NetWeaver (figure 1). To name and document the project, click the properties button
from the knowledge base toolbar. This creates a new project within NetWeaver (figure 1). To name and document the project, click the properties button  on the knowledge base toolbar to open the properties editor for the knowledge base (figure 2). Under Knowledge Base Name enter “Cohead: A NetWeaver2 Tutorial” (figure 3). Click the apply button
on the knowledge base toolbar to open the properties editor for the knowledge base (figure 2). Under Knowledge Base Name enter “Cohead: A NetWeaver2 Tutorial” (figure 3). Click the apply button  . Close this window.
. Close this window.
To begin to build the first dependency network, click on the Create a new dependency network button  on the knowledge base (i.e. topmost toolbar) NetWeaver toolbar. Each newly created topic in NetWeaver has a default name of _Untitledxx. Select this name with your cursor and rename the dependency network “Cohead Habitat” (figure 4)
on the knowledge base (i.e. topmost toolbar) NetWeaver toolbar. Each newly created topic in NetWeaver has a default name of _Untitledxx. Select this name with your cursor and rename the dependency network “Cohead Habitat” (figure 4)
As is the convention for dependency networks, there is an OR node from which we can begin to develop the network. This OR represents the possibility of more than one solution. To hang objects from the OR node, select the node by single clicking it once. Remember that to hang objects from a node, it is necessary to select the node first.
Because experts tell us that we need both spawning habitat and juvenile rearing habitat to define good overall Cohead habitat, it is necessary to AND them. From the network editing toolbar (i.e. the bottom-most toolbar) click the AND button  (figure 5). Now select this newly created AND node to attach objects that represent Cohead spawning habitat and juvenile rearing habitat.
(figure 5). Now select this newly created AND node to attach objects that represent Cohead spawning habitat and juvenile rearing habitat.
To create the Cohead spawning habitat and juvenile rearing habitat networks (remember, first make sure the AND node is selected) click on the drop down menu for adding dependency networks 
 to the existing network on the network editing toolbar (i.e. the bottom-most toolbar). This will result in a network that looks like figure 6. It is necessary to change the names of these new networks from the default _Untitled1 and _Untitled 2. To name a network, double click its oval, select the default name in the network editor and write “Cohead Spawning”. Repeat for the other network to rename it Juvenile Rearing. You should now have a network that looks like figure 7. We have now graphically shown that Cohead habitat quality depends on both Cohead Spawning and Juvenile Rearing. The oval shaped objects that represent these new networks indicate that Cohead spawning and juvenile rearing are, in turn, dependency networks and dependent on other factors.
to the existing network on the network editing toolbar (i.e. the bottom-most toolbar). This will result in a network that looks like figure 6. It is necessary to change the names of these new networks from the default _Untitled1 and _Untitled 2. To name a network, double click its oval, select the default name in the network editor and write “Cohead Spawning”. Repeat for the other network to rename it Juvenile Rearing. You should now have a network that looks like figure 7. We have now graphically shown that Cohead habitat quality depends on both Cohead Spawning and Juvenile Rearing. The oval shaped objects that represent these new networks indicate that Cohead spawning and juvenile rearing are, in turn, dependency networks and dependent on other factors.
Data Links in Dependency Networks
As we have seen, dependency networks can contain dependency networks within themselves. The simple network that we have constructed for Cohead Habitat is an example of the initial top-down approach to knowledge base development. Although we have made no provisions to input data into this simple model, we have created the framework within which we can expand our model to address and interpret data.
Data links are the NetWeaver objects that retrieve, store, and interpret data. Data links compare data values to arguments, and calculate a TRUEness value based upon this comparison ranging from –1 to 1. If, for example, the argument in a data link is “> 3” and the data value is 2, this generates –1 (i.e., FALSE). If the data value is 5, this generates 1 (i.e., TRUE). Therefore, using a fuzzy argument permits the data link to generate TRUEness values between –1 and 1.
Simple Arguments vs Fuzzy Arguments
Experts tell us that Cohead spawning requires good gravel size and low silt content in the gravel.
The best gravel is very fine, ≤1.5 inches in diameter, fair gravel is from 1.5 to 3 inches and unsuitable (bad) gravel is ≥3 inches. Ideally, spawning gravel contains no silt. Good silt content is ≤3 percent, fair silt content ranges from 3 to 7 percent and unsuitable silt content is ≥7 percent (Table 1).
| GOOD | FAIR | BAD | |
|---|---|---|---|
| Gravel Size | ≤ 1.5 | > 1.5 and < 3 | ≥ 3 |
| Silt | ≤ 3 | > 3 and < 7 | ≥ 7 |
Table 1. The argument list for gravel size and silt content relative to Cohead spawning habitat quality.
To better understand the use of data links and arguments within data links, we will use the above information to build two models to describe Cohead Spawning. The first model will use only simple arguments to construct networks defining good, fair and bad spawning habitat quality. The second model will rely on fuzzy arguments to describe Cohead spawning habitat quality.
Simple Argument for Good Spawning Habitat
Task: Build a dependency network depicting good spawning habitat using a simple argument.
Close all windows except the knowledge base project window (figure 1). Create a new dependency network by clicking the new dependency network button from the topmost knowledge base toolbar  . Name the dependency network “Good Spawning”.
. Name the dependency network “Good Spawning”.
For the stream to have good spawning habitat, it needs both good gravel and good silt percentages, therefore hang an AND node from the OR in the network editor by selecting the OR node. Next, click on the AND button  on the dependency network editor toolbar (figure 8) Now we will hang our first data link from the AND node of this network. Be sure the AND node is selected. From the dependency network editor toolbar, locate and click on the drop down menu to the right of the data link button
on the dependency network editor toolbar (figure 8) Now we will hang our first data link from the AND node of this network. Be sure the AND node is selected. From the dependency network editor toolbar, locate and click on the drop down menu to the right of the data link button 
 . From the drop down menu, choose New. This brings up the properties window for the data link where you can name this data link “Gravel”. In the hints field, type a question that asks about the spawning gravel condition. This question might be “What is the average size of gravel in the spawning beds?” (figure 9). Click the accept button
. From the drop down menu, choose New. This brings up the properties window for the data link where you can name this data link “Gravel”. In the hints field, type a question that asks about the spawning gravel condition. This question might be “What is the average size of gravel in the spawning beds?” (figure 9). Click the accept button  to accept the changes.
to accept the changes.
After completing the Data Link Documentation, an argument list window will appear. This window is where one creates arguments for use within a data link. Click the simple argument button 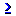 to create a new simple argument (figure 10). This will open the crisp argument definition window (crisp argument is synonymous with simple argument) (figure 11)
to create a new simple argument (figure 10). This will open the crisp argument definition window (crisp argument is synonymous with simple argument) (figure 11)
Remember from Table 1 that good gravel conditions occur when average gravel size is ≤ 1.5 inches. In order to create this argument, one first has to choose the operator ≤ by using the drop down menu under Operator (figure 12). Then the value of 1.5 needs to be entered in the Operand field (figure 13). Click the accept button  in the lower right of the window to accept the new argument. Close the argument viewer window to return to the argument list window for Gravel (figure 14). Close the Gravel arguments window to return to the Good Spawning dependency network editor window. Note that the shape of a data link is a square with the data link name and argument within it.
in the lower right of the window to accept the new argument. Close the argument viewer window to return to the argument list window for Gravel (figure 14). Close the Gravel arguments window to return to the Good Spawning dependency network editor window. Note that the shape of a data link is a square with the data link name and argument within it.
Repeat this procedure by creating another data link to hang from the AND node in the dependency network “Good Spawning”. Name this simple data link “Silt”, and write the question in the comment window: “What is the percent silt in the spawning gravel?”. Follow the previous steps to create the simple argument “≤ 3”. When done, the Good Spawning network is complete (figure 15). Close the Good Spawning dependency network editor.

Simple Argument for Bad Spawning Habitat
Task: Build a dependency network depicting bad spawning habitat using a simple argument.
Repeat the procedure just used to build Good Spawning to create a new Bad Spawning dependency network to define bad spawning habitat. This time, knowing that either bad silt conditions or bad gravel size will prevent Cohead spawning, one does not need to hang an AND from the OR node. You will just create data links to hang directly on the OR node.
You do not need to create new data links for silt and gravel, since you can reuse the data links you created for the Good Spawning network. However, you must create a different argument to define bad spawning. In the data link Gravel create a new crisp argument of ≥ 3. To do this, we are going to use a slightly different method. On the dependency network editor toolbar, rather than using the drop down menu next to the data link button 
 , this time click the button. This will open a list on the right of the main screen showing available topics (in this case data links) for use in the dependency network Bad Spawning (figure 16).
, this time click the button. This will open a list on the right of the main screen showing available topics (in this case data links) for use in the dependency network Bad Spawning (figure 16).
In the available topics list, click Gravel. This brings up the argument list for Gravel. At present, this list consists only of the single argument ≤ 1.5 that we previously built. As before, create a new argument appropriate to defining Bad Spawning (i.e. ≥ 3). After creating this argument, it will appear in the argument list window under both the Arguments used here list and in the right hand list where all arguments for Gravel are listed. Finish with this argument, then return to the list showing available topics. Click on Silt and repeat the above process to create an argument appropriate for Bad Spawning (i.e. ≥ 7). After finishing this argument, the Bad Spawning dependency network should look like figure 17.

Simple Argument for Fair Spawning Habitat
Task: Build a dependency network depicting fair spawning habitat using a simple data link.
Repeat the above process to create a new dependency network named “Fair Spawning” that will define those spawning habitat conditions that lie between good and bad. Give this some thought, as you will have to accommodate all the possible combinations of data that can lead to describing Cohead spawning habitat as fair. If both gravel and silt fall within the fair range, the spawning habitat will be fair. In addition, if the gravel size is good, but the silt is fair (or vice versa) the habitat will also be fair. (Remember that if either gravel size or silt is bad, the spawning habitat will be bad.) This gives us three combinations of silt and gravel that define fair spawning habitat (fair-fair, good-fair and fair-good). If any one of these three conditions is true, then we can say that the spawning habitat is fair (the Fair Spawning dependency network evaluates to TRUE).
From the OR node, hang an AND as we did for Good Spawning. To the AND node attach the data links Gravel and Silt. Remember, these data links are available by using the drop down menu next to the data link button on the dependency network editor toolbar 
 , or by clicking the button itself to bring up the topics selector window. Since the topics selector window is still open, click Gravel to open its arguments list. Define the fair range of values for the arguments (i.e., > 1.5 and < 3 for Gravel; > 3 and < 7 for Silt). Since these are two part arguments (i.e. they contain two operators and two operands, a two step process is needed. Create the first half of the gravel argument like you did above. This argument will read > 1.5. Next, click on the add argument button
, or by clicking the button itself to bring up the topics selector window. Since the topics selector window is still open, click Gravel to open its arguments list. Define the fair range of values for the arguments (i.e., > 1.5 and < 3 for Gravel; > 3 and < 7 for Silt). Since these are two part arguments (i.e. they contain two operators and two operands, a two step process is needed. Create the first half of the gravel argument like you did above. This argument will read > 1.5. Next, click on the add argument button  to add the second operator and operand (i.e. < 3). After completing this step, the argument list should look like figure 18.
to add the second operator and operand (i.e. < 3). After completing this step, the argument list should look like figure 18.
Close the argument list for Gravel and repeat the process for Silt. This will be very similar to what you just did for Gravel, only in this case, the argument will be > 3 and < 7. After completing the arguments for Silt, the Fair Spawning network should look like figure 19.
This network is not finished. As we mentioned earlier, Silt can be fair and Gravel good, or Gravel can be fair and Silt can be good. From the OR node, hang a second AND. From this AND describe and hang Gravel with the fair argument and Silt with the good argument. You can select these already existing arguments from the arguments list for Gravel or Silt. Move the argument you want to use from the list on the right to the left list (Arguments used Here) by selecting the argument and then clicking the move left button  .
.
From the OR node, hang a third AND. From this AND describe and hang Gravel with the good argument and Silt with the fair argument. You have now defined the dependency network describing Fair Spawning for Cohead salmon (figure 20).
Notice that you now have explicitly represented every combination of potential data states in the three networks that you have built for Cohead spawning (i.e., Good Spawning, Fair Spawning, and Bad Spawning). When using simple arguments, it is necessary to do this. Failing to represent all data state combinations can lead to errors in interpreting data.
Fuzzy Arguments for Spawning Habitat
Task: Build a dependency network for Cohead spawning using fuzzy arguments.
Again follow the dependency network creation process to create a new network named “Fuzzy Spawning”. The intent of this network is to characterize optimal Cohead spawning habitat. Again, because both Silt and Gravel must be optimal for spawning habitat to be optimal, hang an AND node from the OR node. To the AND node attach the Gravel data link. This time, create a fuzzy argument by clicking the fuzzy argument button  in the argument list window (figure 18). Name this argument “Fuzzy Gravel”. Think of the fuzzy argument as a curve, with the data values (e.g., gravel size) as the x axis and TRUEness (or, in this case, goodness of the habitat) as the y-axis. For Gravel, enter 1.5 in the field labeled x. In the y field, enter 1 (for true) or click the true button
in the argument list window (figure 18). Name this argument “Fuzzy Gravel”. Think of the fuzzy argument as a curve, with the data values (e.g., gravel size) as the x axis and TRUEness (or, in this case, goodness of the habitat) as the y-axis. For Gravel, enter 1.5 in the field labeled x. In the y field, enter 1 (for true) or click the true button  to enter the 1. This indicates that values ≤ 1.5 are TRUE (i.e., good gravel size). Repeat this by entering 3 in the next open field labeled x and entering a -1 (for false) or click the false button
to enter the 1. This indicates that values ≤ 1.5 are TRUE (i.e., good gravel size). Repeat this by entering 3 in the next open field labeled x and entering a -1 (for false) or click the false button  to enter the -1.
to enter the -1.
Now, view your new fuzzy argument by clicking the try button 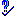 . This will display the fuzzy curve for this argument in the left side pane. The fuzzy argument should now range from 1.5 (TRUE) to 3.0 (FALSE) (figure 21). Click on the accept button
. This will display the fuzzy curve for this argument in the left side pane. The fuzzy argument should now range from 1.5 (TRUE) to 3.0 (FALSE) (figure 21). Click on the accept button  to accept the new argument. This displays the argument in a window that should now be closed.
to accept the new argument. This displays the argument in a window that should now be closed.
Repeat this process by attaching the Silt data link to the AND node in the Fuzzy Spawning dependency network. Name the fuzzy argument “Fuzzy Silt”. It should range from 3 (TRUE) to 7 (FALSE) (figure 22). Now, you have created a dependency network describing Cohead spawning habitat quality using fuzzy arguments (figure 23). Compare this network to the networks required to describe spawning habitat quality using simple arguments (figures 17, 19 & 20).
Testing a Knowledge Base
We have now built two different models for defining Cohead spawning habitat quality. In the first model, we used simple arguments to create three dependency networks that define good, fair, and bad spawning habitat qualities. In the second model, we used fuzzy arguments to build a single dependency network to define spawning habitat. We will now test these different approaches and compare the utility of the two models.
One way to run/test a knowledge base is to build a goal group (aka dependency network group) containing the dependency networks that you want to evaluate. The group is a useful tool for comparing the performance of multiple dependency networks at the same time. There are no restrictions on which dependency networks you can place in a group, but as a general guideline, a group typically contains related dependency networks such as those just built to describe Cohead spawning habitats.
Task: Build a group of goal topics to test your knowledge base.
From the knowledge base toolbar, click the goal group button  . This brings up a window containing two panes and an upper field for naming the group. Name this group “Cohead Spawning Group”. In the right pane (labeled Knowledge Base) are all the dependency networks in the knowledge base. In the left pane, you will place those dependency networks that you want to include in this group. Like the arguments list window where you can select an argument from the right list and move them to the left using the move left arrow button
. This brings up a window containing two panes and an upper field for naming the group. Name this group “Cohead Spawning Group”. In the right pane (labeled Knowledge Base) are all the dependency networks in the knowledge base. In the left pane, you will place those dependency networks that you want to include in this group. Like the arguments list window where you can select an argument from the right list and move them to the left using the move left arrow button  , the same procedure is used for adding dependency networks to the group list. Add Good Spawning, Fair Spawning, Bad Spawning, and Fuzzy Spawning to the group (figure 24).
, the same procedure is used for adding dependency networks to the group list. Add Good Spawning, Fair Spawning, Bad Spawning, and Fuzzy Spawning to the group (figure 24).
Close the Cohead Spawning Group window. From the knowledge base project window you will now see that the Cohead Spawning Group is now listed under the outline tab  . Double-click Cohead Spawning Group. This will open the interrogation window for this group (figure 25). At the bottom of this window you will see two panes. The Unresolved Influential Topics pane lists all topics for which we have provided no data. The Other Antecedent Topics pane lists all those topics that have been provided data. This list is empty for now as we have provided no data. Above these two panes is a box in which one can add data values, or select data values from a drop down menu. Above this is the name and question for the current topic.
. Double-click Cohead Spawning Group. This will open the interrogation window for this group (figure 25). At the bottom of this window you will see two panes. The Unresolved Influential Topics pane lists all topics for which we have provided no data. The Other Antecedent Topics pane lists all those topics that have been provided data. This list is empty for now as we have provided no data. Above these two panes is a box in which one can add data values, or select data values from a drop down menu. Above this is the name and question for the current topic.
Now we are ready to enter some data. When we created each data link, we entered questions in the Hints field. As each question appears (e.g., “What is the average size of gravel in the spawning beds?”), provide the value in the data entry field. Once the data value is entered it is automatically applied to the knowledge base. For this example, we will enter 2 for gravel and 4 for silt. As we enter data for each topic, the topic will shift to the right pane. Only the unresolved topics will stay in the left pane.
After we have entered all the required data, the Cohead Spawning Group window will display the results of the interrogation (figure 25). The more TRUE a dependency network is the more its TRUEness bar extends to the right and becomes greener. The more FALSE a dependency network is the more its TRUEness bar extends to the left and the redder it becomes.
Notice that, after entering values for gravel size and silt percentage, only one of the three networks from the simple argument model (i.e., Good Spawning, Fair Spawning, and Bad Spawning) is fully TRUE, and the other two networks are fully FALSE (i.e., not TRUE). What about fair values at the bad and good ends of the spectrum? How can you know that you have a low fair or a high fair?
Note that:
- Fuzzy Spawning is fully TRUE whenever Good Spawning is fully TRUE.
- Fuzzy Spawning is fully FALSE whenever Bad Spawning is fully TRUE.
- Fuzzy Spawning assumes intermediate values whenever Fair Spawning is fully TRUE.
In the Cohead Spawning Group window, click to select Fair Spawning and right click to bring up a context sensitive menu. Select view/run… . This will bring up the Fair Spawning dependency network window with colors indicating the relative trueness of all the components of the network (figure 26). As you can see, the colors here represent that Fair Spawning is true because gravel is between 1.5 and 3 (i.e. 2) and silt is between 3 and 7 (i.e. 4). It is only necessary for one path to the OR node to be true for the network to calculate to true. Only if all paths to the OR are false will Fair Spawning calculate to False.
. This will bring up the Fair Spawning dependency network window with colors indicating the relative trueness of all the components of the network (figure 26). As you can see, the colors here represent that Fair Spawning is true because gravel is between 1.5 and 3 (i.e. 2) and silt is between 3 and 7 (i.e. 4). It is only necessary for one path to the OR node to be true for the network to calculate to true. Only if all paths to the OR are false will Fair Spawning calculate to False.
Close Fair Spawning. Using the same procedure used to view/run Fair Spawning, open Fuzzy Spawning (figure 26). As you can see, although the network is green, it is not as vividly green as the true elements of the Fair Spawning network. Mousing over any element of the network will display that elements data value and fuzzy set membership. The uppermost OR is 38.9% True (i.e. .389 on the continuum from -1 to 1.
Fuzzy Spawning (figure 26) is capable of representing the bad fair, fair fair, and good fair descriptions of the habitat in the continuum of values between Good Spawning and Bad Spawning in addition to representing good and bad spawning habitat conditions. Fuzzy Spawning by itself, is capable of rendering the entire Cohead spawning habitat description. Diligent and careful application of fuzzy arguments in data links reduces the knowledge engineering effort since all one needs to represent as dependency networks are fuzzy descriptions of ideal system states.
Editing Dependency Networks
NetWeaver supports normal editing features and these can be found in the network editor toolbar. The delete, cut, copy, and paste functions are applied to the selected node in a dependency network and all nodes hanging from it (aka antecedents or children).
Now that we have created a preliminary, but functional, fuzzy model for Cohead spawning, let’s practice using copy and paste by replacing the incomplete dependency network Cohead Spawning that was in our original knowledge base template (figure 6) with Fuzzy Spawning. This replacement can be done several ways. We will use the two most common approaches, namely: 1) replace one dependency network with another, and 2) paste the contents of one network into another network.
Task: Replace Cohead Spawning with Fuzzy Spawning.
Close all open windows until only the knowledge base project window (figure 1) is open. Click on the dependency networks tab  to display all the dependency networks in the knowledge base (figure 27). Double-click on the Cohead Habitat dependency network to open its dependency network window (figure 3).
to display all the dependency networks in the knowledge base (figure 27). Double-click on the Cohead Habitat dependency network to open its dependency network window (figure 3).
Select the AND node in the Cohead Habitat dependency network window. From the topic selection window (which should still be open to the right of your screen) the list of data links for the knowledge base are presently displayed. To display a list of dependency networks, use the drop down menu at the top of the topic selection window to select dependency networks  . This displays a list of dependency networks in the knowledge base. You will notice that Cohead Habitat is grayed-out. If an item is grayed out in this list it is because it can not be used due to circular reasoning (i.e. Cohead Spawning cannot contain Cohead Spawning). Click Fuzzy Spawning in this list to hang Fuzzy Spawning from the selected AND node in the Cohead Habitat dependency network (figure 28).
. This displays a list of dependency networks in the knowledge base. You will notice that Cohead Habitat is grayed-out. If an item is grayed out in this list it is because it can not be used due to circular reasoning (i.e. Cohead Spawning cannot contain Cohead Spawning). Click Fuzzy Spawning in this list to hang Fuzzy Spawning from the selected AND node in the Cohead Habitat dependency network (figure 28).
While still in the Cohead Habitat dependency network, select the Cohead Spawning network oval and delete it by pressing the delete key on your keyboard, or by clicking on the delete button  on the dependency network editor toolbar.
on the dependency network editor toolbar.
You have now modified the original Cohead Habitat dependency network so that it connects to a functional model of Cohead spawning quality, i.e., Fuzzy Spawning (figure 29).
Sometimes it is convenient to rearrange nodes within a network. Nodes can be moved by selecting them and moving them left or right using the arrow buttons 
 on the network editing toolbar. For example: suppose we wanted to move Fuzzy Spawning from the right side of the AND node in Cohead Habitat to the left side. To do so, select the oval representing Fuzzy Spawning and then click on the move left button
on the network editing toolbar. For example: suppose we wanted to move Fuzzy Spawning from the right side of the AND node in Cohead Habitat to the left side. To do so, select the oval representing Fuzzy Spawning and then click on the move left button  on the dependency network editor toolbar. This will cause Fuzzy Spawning to move to the left and the network will now look like figure 30.
on the dependency network editor toolbar. This will cause Fuzzy Spawning to move to the left and the network will now look like figure 30.
Another method for replacing one dependency network with another is to copy or cut the contents of one dependency network and paste those contents into another network. Before we demonstrate this, restore the Cohead Spawning dependency network to the Cohead Habitat dependency network and delete Fuzzy Spawning so that the Cohead Habitat dependency network is restored as in figure 6.
There are two paste functions 
 , and you should be careful about which function you use. Paste the copied node…
, and you should be careful about which function you use. Paste the copied node…  pastes the chosen node and everything hanging from it (its antecedents or children). Paste the Children…
pastes the chosen node and everything hanging from it (its antecedents or children). Paste the Children…  pastes only those nodes hanging from the chosen node, but not the node itself.
pastes only those nodes hanging from the chosen node, but not the node itself.
To demonstrate the difference, double click the oval representing Cohead Spawning to open its dependency network window. At the present time, this network contains only an OR node. Go to the Fuzzy Spawning dependency network window (figure 31), and select the topmost OR. From the dependency network editor toolbar, click on the Copy button  . Return to the Cohead Spawning dependency network window. From the toolbar click the Paste the children…. Button . This will result in the contents of the Fuzzy Spawning dependency network being pasted into the Cohead Spawning dependency network (figure 32). By choosing to Paste the children…
. Return to the Cohead Spawning dependency network window. From the toolbar click the Paste the children…. Button . This will result in the contents of the Fuzzy Spawning dependency network being pasted into the Cohead Spawning dependency network (figure 32). By choosing to Paste the children…  you will be pasting all those objects antecedent (i.e., hanging from, the children of the node) to the selected node in Fuzzy Spawning. If you had chosen Paste the copied node…
you will be pasting all those objects antecedent (i.e., hanging from, the children of the node) to the selected node in Fuzzy Spawning. If you had chosen Paste the copied node…  to perform this operation, you would have pasted the topmost OR as well as its antecedents (children), adding an extra, incorrect OR node (figure 33).
to perform this operation, you would have pasted the topmost OR as well as its antecedents (children), adding an extra, incorrect OR node (figure 33).
To address the redundant OR nodes in figure 33, select the lower OR node, and delete it  . This will remove all but the top most OR. On the dependency network editor toolbar, click Paste the children
. This will remove all but the top most OR. On the dependency network editor toolbar, click Paste the children  . This will result in a network looking like figure 32.
. This will result in a network looking like figure 32.
At this time, close all dependency network windows leaving open only the knowledge base project window.
Calculated Data Links
We must often perform mathematical manipulation of data to derive a value to be used in a knowledge base. NetWeaver facilitates this via the calculated data link. Calculated data links are equations built as networks whose nodes are not necessarily Boolean operators (e.g., AND and OR) that are used in dependency networks, but instead are mathematical operators (e.g., + and -). Calculated data links are created at the knowledge base level by selecting the create calculated data link button 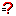 on the knowledge base toolbar, or by using
on the knowledge base toolbar, or by using 
 on the dependency network editor toolbar if you are adding it to an existing, open network. Let’s build a calculated data link within the Cohead knowledge base.
on the dependency network editor toolbar if you are adding it to an existing, open network. Let’s build a calculated data link within the Cohead knowledge base.
For this example, we will build a calculated data link to assist in determining if sufficient spawning gravel exists in a stream segment. We have already dealt with spawning gravel conditions insofar as gravel size and percent silt are concerned. However, even if these parameters are optimal, we may still want to know: “Is there enough of this spawning habitat in a given stream segment for it to be a good spawning habitat?” Our experts tell us that there must be at least 300, and ideally 500 square meters of spawning gravel comprising more than 20 percent, and ideally 30 percent of the stream reach in order to characterize the habitat as optimal for Cohead spawning.
We will build a dependency network to reflect the fact that good Cohead spawning habitat depends on:
- the size of the gravel
- the percent silt in the gravel
- the total area of spawning gravel available
- the percent of the stream bed that is made up of this gravel
Silt percent, gravel size and gravel area are data values so we use simple data links for these values. Percent gravel, however is calculated from two data values, gravel area and total stream reach area, therefore, we need to use a calculated data link.
To build this portion of the knowledge base, open the Cohead Spawning dependency network. This dependency network currently ANDs the data links for silt and gravel, which use the arguments Fuzzy Silt and Fuzzy Gravel, respectively (figure 33). Select the AND node and on the topics selector window (which is still open on the right of your screen) bring up the list of calculated data links using the drop down menu 
 . Since we have no calculated data links as yet, the only choice available is new. Click on new to bring up an untitled arguments list. Click on the fuzzy argument button
. Since we have no calculated data links as yet, the only choice available is new. Click on new to bring up an untitled arguments list. Click on the fuzzy argument button  to create a new fuzzy argument. Name this argument “Fuzzy Percent Gravel”. Create a fuzzy argument where it is FALSE when ≤ 20 and TRUE when ≥ 30. Click on the accept button
to create a new fuzzy argument. Name this argument “Fuzzy Percent Gravel”. Create a fuzzy argument where it is FALSE when ≤ 20 and TRUE when ≥ 30. Click on the accept button  to accept this argument resulting in a fuzzy curve like that shown in figure 34. Close this window and then close the argument list window.
to accept this argument resulting in a fuzzy curve like that shown in figure 34. Close this window and then close the argument list window.
The Cohead Spawning dependency network now has three data links attached to the AND node (figure 35). Double click on the new calculated data link to open its window. Select the _Untitled name and overwrite it with the name “Percent Spawning Gravel”. Now we will build the equation to calculate percent spawning gravel. This equation can be stated as:
Working from the Percent Spawning Gravel calculated data link window (note that the topmost node is an equals sign [=] sign rather than the OR at the top of dependency networks), go to the Functions menu 
 (note how this menu is enabled now that we are working in a calculated data link). This menu is to the far right of the lowermost toolbar. From the drop down menu
(note how this menu is enabled now that we are working in a calculated data link). This menu is to the far right of the lowermost toolbar. From the drop down menu 
 , select the multiplication symbol [*]. This attaches the multiplication symbol to the equals node. Select the multiplication node and, from the functions menu
, select the multiplication symbol [*]. This attaches the multiplication symbol to the equals node. Select the multiplication node and, from the functions menu 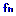
 select the division symbol [/] to attach it to the multiplication node. With the multiplication node still selected, go to the Constants menu
select the division symbol [/] to attach it to the multiplication node. With the multiplication node still selected, go to the Constants menu 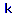
 and select the [k] node. This node allows users to define constants in equations. Select this constant node, and right click for the contextual menu and open its Properties
and select the [k] node. This node allows users to define constants in equations. Select this constant node, and right click for the contextual menu and open its Properties  . In the properties window, enter 100 in the Value/constant name field. Click the apply button
. In the properties window, enter 100 in the Value/constant name field. Click the apply button  to apply this constant. Close the properties window.
to apply this constant. Close the properties window.
Now select the division node. It is to this node that we wish to attach two new data links, one for total area of the stream reach, and the other for area of spawning gravel within the stream reach. To do this, we return to the topics list and change the list to show data links 
 . Select new from this list, and name this new data link “Area of Spawning Gravel”. In the hints field write “What is the area of spawning gravel (sq meters)?”. Click the accept button
. Select new from this list, and name this new data link “Area of Spawning Gravel”. In the hints field write “What is the area of spawning gravel (sq meters)?”. Click the accept button  . Repeat to create another data link. Name this link “Total area of stream reach” and in the hint field write “What is the total area of the stream reach (sq meters)?” and accept it. At this point, your network should look like figure 36.
. Repeat to create another data link. Name this link “Total area of stream reach” and in the hint field write “What is the total area of the stream reach (sq meters)?” and accept it. At this point, your network should look like figure 36.
It is important that the objects hung from a division node are in the correct order. In other words, for any mathematical node such as subtraction or division, the order of the two variables matters. The data link Area of Spawning Gravel is to the left and the data link Total area of stream reach is to the right (figure 36). The network, when read from right to left is the same as the equation above. When the data link is blue, it contains data. When the data link is orange, it does not yet contain data, or it is unable to correctly calculate its value.
Since Cohead populations require both adequate spawning gravel area and adequate gravel as a percent of the stream reach area, we will create a separate fuzzy argument for gravel area. Close the Percent Spawning Gravel calculated data link window. In the Cohead Spawning dependency network window select the AND node. To this AND node, hang the data link Area of Spawning Gravel, and create a new fuzzy argument named “Fuzzy Gravel Area” that ranges from 300 (FALSE) to 500 (TRUE) (figure 37). Accept this argument, review the shape of the argument in the fuzzy argument window and close it. Then close the argument list window. Now the Cohead Spawning dependency network is complete (Figure 38). Note that the Area of spawning gravel data values are used twice in the Cohead spawning dependency network: once in the Fuzzy Gravel Area argument, and once as part of the Percent Spawning Gravel calculated data link.
Task: Create a temporary goal group to test the Cohead Spawning dependency network.
When we compared the simple and fuzzy argument approaches for Cohead spawning, we created a new Goal Group (figure 19). A similar way to test the Cohead Spawning dependency network would be to create a new group by clicking on the Create a new goal group topic button  on the knowledge base toolbar.
on the knowledge base toolbar.
A second way to test networks is to create a temporary group. This is useful when there is only one dependency network or calculated data link that we want to test. We can build a temporary group by right clicking within the Cohead Spawning dependency network window (don’t click on a node but rather click in white space). From the contextual menu, choose Open as goal group  . This will create a temporary goal group consisting only of the dependency network or calculated data link that you want to test. In the group window that is now open, provide the following data:
. This will create a temporary goal group consisting only of the dependency network or calculated data link that you want to test. In the group window that is now open, provide the following data:
- Silt: 3
- Gravel: 2
- Area of Spawning Gravel: 450
- Total Area of Stream Reach: 550.
The group window should now look like figure 38.
Fuzzy Curves
Now that we have completed the Cohead Spawning component of the Cohead Habitat knowledge base, we will turn our attention to Juvenile Rearing. For Cohead salmon, the main requirement for suitable habitat for rearing juveniles is adequate areas of deep pools. The expected percentage of the surface area of a stream segment (i.e., reach) that is over deep pools changes with the steepness (i.e., gradient) of the stream, or reach gradient. The steeper the reach gradient, the lower the area of deep pool one can expect. The optimum percentage of deep pool required by the juvenile salmon also changes with the reach gradient, and there can be too little pool or too much pool for the young to survive.
The conventional method for appraising the pool habitat for Cohead juvenile rearing is to classify the stream reach gradient into three categories (i.e., ≤ 2%, > 2% and < 5%, and ≥ 5%). For each of these gradient categories, we need to construct fuzzy arguments that portray the ideal expectation for percent pool. The fuzzy percent pool arguments are two tailed (i.e., there can be too little as well as too much pool for a given gradient), unlike the one tailed arguments we used earlier for Percent Silt and “Gravel Size.
To construct the Juvenile Rearing dependency network, open its dependency network window. It should only contain an OR node. To this node, attach an AND node. Select the AND node and attach a new data link and name it Reach Gradient. In the hint field write: “What is the gradient of the stream reach (percent)?”. Create the simple (crisp) argument of ≤ 2. Accept this argument and close windows until you are looking at the Juvenile Rearing dependency network window. Select the AND node and attach another new data link. Name this data link Percent Pool and in the hint field write: “What is the percentage of pools in the stream reach?”. Build a fuzzy argument where 15 is FALSE, 55 is TRUE, 70 is TRUE, and 85 is FALSE. Name this argument “Gradient ≤ 2”. Accept this argument. Your argument should look like figure 39.
In the Juvenile Rearing dependency network window, select the uppermost OR and hang another AND node to it. To this AND node attach the data link Reach Gradient and create a new crisp argument > 2 and < 5. Accept the argument and close the various windows until you are once again looking at the Juvenile Rearing dependency network. As we did on the first AND node, attach the data link Percent Pool to the second AND. Create a fuzzy argument in Percent Pool of 10 is FALSE, 40 is TRUE, 60 is TRUE, and 80 is FALSE. Name this argument “Gradient > 2 and < 5”.
Close windows and return to the Juvenile Rearing dependency network. Add another AND to the topmost OR. To this AND hang the Reach Gradient data link and create a new crisp argument of ≥ 5. Accept the argument and add the Percent Pool data link to this third AND. In the Percent Pool data link, create a fuzzy argument of 0 is FALSE, 30 is TRUE, 50 is TRUE, and 70 is False. Name this argument “Gradient ≥ 5”. The Juvenile Rearing dependency network should now look like figure 40.
Although the Percent Pool data links use fuzzy arguments, the Reach Gradient data links use simple arguments. As a consequence, the transition between interpretations of appropriate percent pool for each reach gradient category (i.e., bad, fair, good) is arbitrary and abrupt. NetWeaver offers a more sensitive and accurate means to characterize relationships such as this. Typically, these relationships can be represented as those in which a question must be answered by a question. In this example, the question, “Is the percent pool on the stream reach good for Cohead juvenile rearing?” would be answered by the question, “What is the reach gradient?”
Task: Build the Fuzzy Curve Juvenile Rearing dependency network using a fuzzy node.
To make room for this exercise, close the topics list window on the right of your screen if it is still open. Create a new dependency network and call it “Fuzzy Curve Juvenile Rearing”. From the network editor toolbar, click on the Create a fuzzy curve drop down menu 
 . In the menu that appears, choose Data Link…
. In the menu that appears, choose Data Link…  . From the data link menu choose Percent Pool. This will hang a fuzzy curve controlled by the data link Percent Pool from the topmost OR node. Select the Percent Pool fuzzy curve node and click on the Create a fuzzy curve button
. From the data link menu choose Percent Pool. This will hang a fuzzy curve controlled by the data link Percent Pool from the topmost OR node. Select the Percent Pool fuzzy curve node and click on the Create a fuzzy curve button  on the network editor toolbar. This will hang an x,y pair node from the fuzzy curve object. Repeat this step three times until there are four x,y pairs hanging from the Percent Pool fuzzy curve (figure 41).
on the network editor toolbar. This will hang an x,y pair node from the fuzzy curve object. Repeat this step three times until there are four x,y pairs hanging from the Percent Pool fuzzy curve (figure 41).
We will use each of these x,y pairs to define the break points in the fuzzy curve for Percent Pool. Break points are points of inflection in a fuzzy curve (places where the slope of the fuzzy curve changes). Remember that the three fuzzy arguments we built for percent pool all had four points. In this case, the left most point is the data value below which the argument is FALSE, the middle two points define the range in which the argument is TRUE, and the right most point defines the date value above which the argument is FALSE. In Fuzzy Juvenile Rearing, the four x,y pairs represent these points. We will now build on these x,y pairs so that the interpretation of Percent Pool makes the transition smoothly from the ≤ 2% reach gradient to the ≥ 5% gradient interpretations.
We start with the leftmost of the four x,y pairs. This will determine the pool percent below which the argument is FALSE (i.e., the pool percent is not enough for successful juvenile rearing) at a given reach gradient. Select the x,y pair node and hang from it another fuzzy curve based upon the data link Reach Gradient. Now hang the constant FALSE from the drop down menu for constants 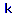
 . (figure 42).
. (figure 42).
Repeat this process of attaching a fuzzy curve based on Percent Pool to the other three x,y pairs. To the second and third x,y pairs, hang the constant TRUE. To the fourth x,y pair hang the constant False. The network should look like figure 43.
We now turn our attention to completing construction of the rest of the Reach Gradient fuzzy curves. To each of the four Reach Gradient fuzzy curves attach two x,y pairs (figure 44).
For the leftmost Reach Gradient fuzzy curve, hang two constant nodes to each x,y pair. Assign values of 2, 15, 5, and 0 respectively (i.e. from left to right)(figure 45).
Repeat for the next Reach Gradient fuzzy curve and assign the constant values of 2, 55, 5, and 30. Repeat again for the third Reach Gradient fuzzy curve and assign the constant values of 2, 70, 5, and 50. Repeat once more for the fourth Reach Gradient fuzzy curve and assign the constant values of 2, 85, 5, and 70 (figure 46).
This network can be interpreted as follows: For the leftmost Reach Gradient fuzzy curve we can interpret this structure as follows: When Reach Gradient values are 2% or less, the left most break point for the Percent Pool fuzzy argument, below which all values are FALSE, is 15. For “Reach Gradient” values of 5% or greater, this left most break point is 0. For Reach Gradients between 2% and 5%, this left most FALSE break point scales between 15 and 0.
The Reach Gradient Fuzzy Curve nodes calculate the transition points on the curve that will determine the optimal percent pool at a given reach gradient. The Percent Pool Fuzzy Curve node, in turn, calculates the TRUEness of the argument based on the transition points calculated by the Reach Gradient Fuzzy Curve nodes. This Percent Pool Fuzzy Curve network is a more inclusive and sensitive representation of the Juvenile Rearing dependency network than depicted in figure 40.
We have now constructed dependency networks that describe the requirements for Cohead spawning and juvenile rearing. We started with a broad description of the Cohead salmon’s habitat requirements, divided the requirements into individual dependency networks, and “built down” until we could describe the requirements as data values. We can now combine these networks with a single dependency network to determine the quality of the Cohead habitat given the following data: average gravel size, percent silt in the gravel beds, area of spawning gravel, total area of stream reach, reach gradient, and percentage of the surface area of the reach that is over deep pools.
